To illustrate this process, I will show a starting Excel file and a much better example.
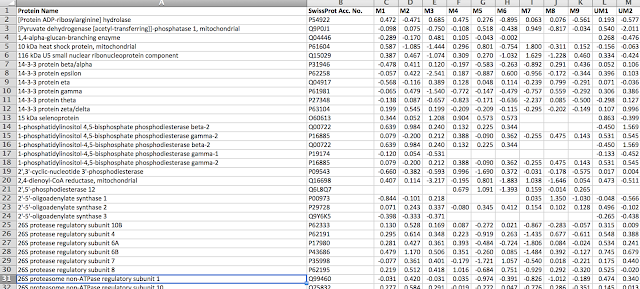
The before file is one that we published last year (Proteomics-Based Strategies To Identify Proteins Relevant to Chronic Lymphocytic Leukemia (Alsagaby et al, J. Proteome Res., 2014, 13 (11), pp 5051–5062). Please download and open our list of 728 proteins which is available from the JPR website as an Excel file.
Here is a photo of the file:
To use this in R:
- Delete everything you don't want.
- Get the titles into one line
- Shorten the titles to something useful
- Save as a csv file (comma separated values)
- Try the read.csv() function to get it into R.
Here is a nicer look file:
There is a way to clean up your data in R too. I used the readxl package.
For an example see this script:





No comments:
Post a Comment
Comments and suggestions are welcome.