Here is a script to do a protein assay:
# Protein Concentrations
prot <- c(0.000, 0.016, 0.031, 0.063, 0.125, 0.250, 0.500, 1.000, 0.000, 0.016, 0.031, 0.063, 0.125, 0.250, 0.500, 1.000)
# Absorbance from my protein assay
abs <- c(0.329, 0.352, 0.349, 0.379, 0.417, 0.491, 0.668, 0.956, 0.327, 0.341, 0.355, 0.383, 0.417, 0.446, 0.655, 0.905)
#Plot the data simply
plot(abs~prot)
#Calculate the line using the linear model function
line <- lm(abs~prot)
#Draw the line
abline(line)
#Improve the graph:
plot(abs~prot,
xlab = "[Protein] (microg/ml)",
ylab = "Absorbance (570nm)",
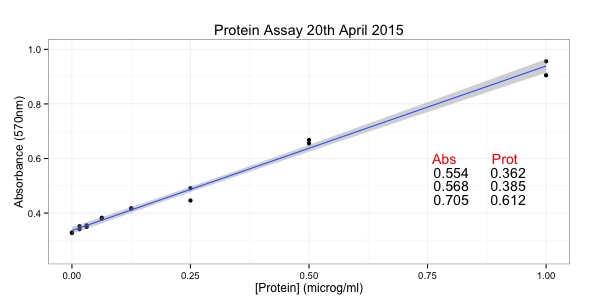
main = "Protein Assay 20th April 2015")
abline(line)
r2 <- round(summary(line)$r.squared, 3)
mylabel = bquote(italic(R)^2 == .(format(r2, digits = 3)))
text(x = 0.2, y = 0.9, labels = mylabel)
#Equation of a line y = mx + c
#In our case abs = slope * prot + intercept
# ukn.prot = (abs - intercept)/slope
int <- summary(line)$coefficients[1]
slope <- summary(line)$coefficients[2]
mylabel = bquote(y == .(format(slope, digits = 3))*x + .(format(int, digits = 3)))
text(x = 0.2, y = 0.8, labels = mylabel)
#now calculate some unknown protein concs from absorbances
#put the unknowns into a vector
abs.unknowns <- c(0.554, 0.568, 0.705)
#rearrange the equation of the line to ukn.prot = (abs - intercept)/slope
prot.unknowns <- (abs.unknowns - int)/slope
#put the answers on the graph
text(x = 0.8, y = (0.6), "Abs")
text(x = 0.92, y = (0.6), "Prot")
for (i in 1:length(abs.unknowns)){
text(x = 0.8, y = (0.6 - i/20), abs.unknowns[i])
text(x = 0.92, y = (0.6 - i/20), round(prot.unknowns[i], 3))
}
# END OF SCRIPT
This can be copy and pasted into the script window of R-Studio.
Then each line can be run one by one by pressing the Run button:
This will generate the following output:




No comments:
Post a Comment
Comments and suggestions are welcome.